parcats 0.0.1 released
parcats was released on CRAN. It is an htmlwidget providing bindings to the plotly.js parcats trace, which is not supported by the plotly R package. Also adds marginal histograms for numerical variables.
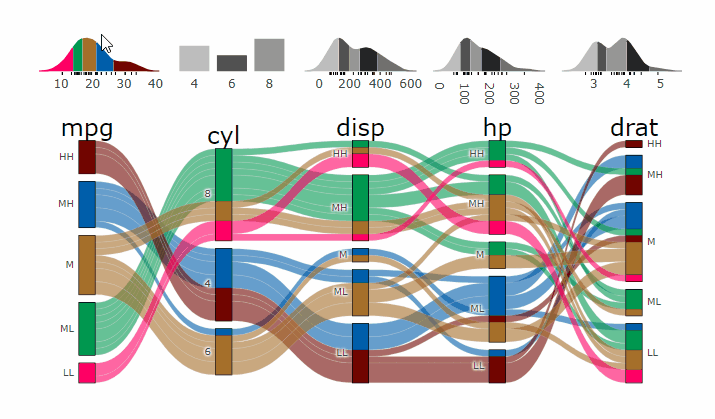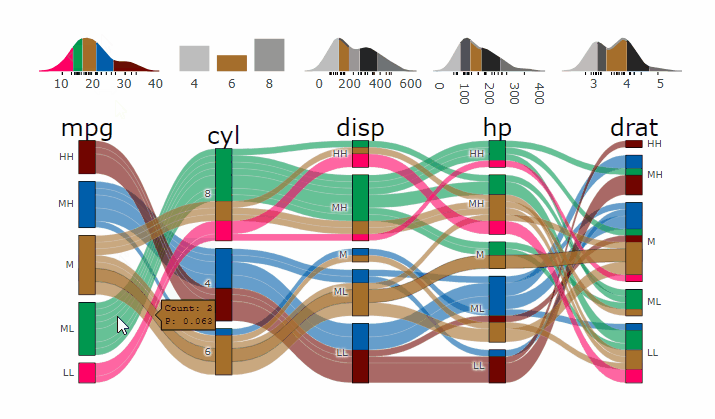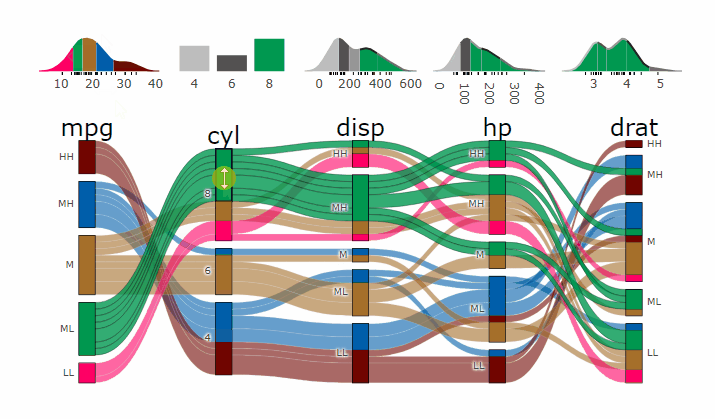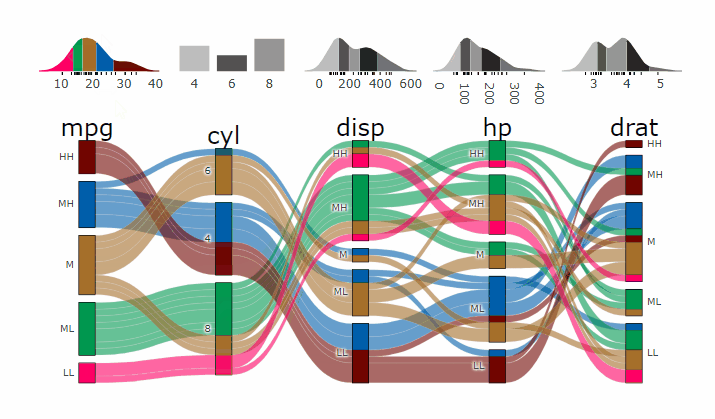
demogif
What it can do
I wanted to add interactivity to easyalluvial plots for a while now and found that the parcats trace of plotly.js would be perfect because brushing with the mouse highlights the entire flow and not just everything flowing in and out of a specific node as in most D3 Sankey chart implementations. Unfortunately the parcats trace was not available in the plotly R package so I decided to build a new html widget to create R bindings for specifically this trace.
- converts any
easyalluvialplot to an interactive parallel categories diagram - interactive marginal histograms
- multidimensional partial dependency and model response plots
easyalluvial
parcats requires an alluvial plot created with easyalluvial to create an interactive parrallel categories diagram.
Demo
Examples
suppressPackageStartupMessages( require(tidyverse) )
suppressPackageStartupMessages( require(easyalluvial) )
suppressPackageStartupMessages( require(parcats) )Parcats from alluvial from data in wide format
p = alluvial_wide(mtcars2, max_variables = 5)
parcats(p, marginal_histograms = TRUE, data_input = mtcars2)Parcats from model response alluvial
Machine Learning models operate in a multidimensional space and their response is hard to visualise. Model response and partial dependency plots attempt to visualise ML models in a two dimensional space. Using alluvial plots or parrallel categories diagrams we can increase the number of dimensions.
Here we see the response of a random forest model if we vary the three variables with the highest importance while keeping all other features at their median/mode value.
df = select(mtcars2, -ids )
m = randomForest::randomForest( disp ~ ., df)
imp = m$importance
dspace = get_data_space(df, imp, degree = 3)
pred = predict(m, newdata = dspace)
p = alluvial_model_response(pred, dspace, imp, degree = 3)
parcats(p, marginal_histograms = TRUE, imp = TRUE, data_input = df)