Use jupyter notebooks to add posts to hugo blog
Table of Contents
The file paths mentioned in this post only apply when the following configuration is set in your ‘.toml’ file:
[permalinks]
post = "/:year/:month/:slug/"
hugo_jupyter
Starting this blog with blogdown and RStudio was pretty straight forward. I did not need to learn much about hugo which is the static website generator that I used and the project is well-documented. However I would also like to add posts from juypter notebooks which seems to be possible but let’s say less accessible. The tool I tried is hugo_jupyter which is supposed to work similar to blogdown. It runs a local server of the website which automatically rerenders the site if changes are made in relevant folders. The same can also be achieved if executing this simple hugo command inside the parent folder of your site (You have to open your browser and copy paste the displayed local host address though).
hugo server
hugo_jupyter will additionally watch a ./notebooks for changes in any jupyter notebooks it contains and convert them into .md to ./content/posts. For this you have to manually add the front-matter parameters title, date, slug, subtitle to the json metadata of your notebook using the jupyter GUI. Any other parameter that you add will not be passed to the .md file of your post. Any graphical output produced by any of your cells will be dropped and replaced by a link in the markdown format . Then hugo will render your .md to .html automatically adding to the image path it finds in the markdown synthax like this yyyy/mm/slug/image_path. In order for this link to work we would need to place the image in ./static/yyyy/mm/slug.
All in all this would require a lot of manual steps after conversion of the notebook to .md format that I decided to only use the code of hugo_jupyter that converts the notebook to .md and make some changes to it so it also extracts images and places them in the appropriate ./static/yyyy/mm/slug path.
| Feature | blogdown | hugo_jupyter |
|---|---|---|
| serve locally | yes | yes |
| encode front-matter parameters | yes | only 4 |
| render graphical code output | yes | no |
| shortcode | as function | yes |
| markdown support | pandoc | hugo |
(table generated with https://www.tablesgenerator.com/)
Modified workflow
- I use the
hugo servercommand in the terminal - I added a
.pythonfolder with the render_notebooks.py file that contains the code added below which I modified fromhugo_jupyterto (i) render all output plots and put the.pngfiles in the appropriate paths and (ii) add front-matter parameters by passing them as kwargs to the render function. - I can view all changes on my localhost server.
- Upload changes to github
- Netlify is linked to my github and automatically renders the whole page
from nbconvert import MarkdownExporter
from nbconvert.preprocessors import Preprocessor
from pathlib import Path
from traitlets.config import Config
import nbformat
import re
path = r'./notebooks/adding_posts_with_juypter.ipynb'
# here I customize some functions of the fabfile.py of the hugo_jupyter package
class CustomPreprocessor(Preprocessor):
"""Remove blank code cells and unnecessary whitespace."""2
def preprocess(self, nb, resources):
"""
Remove blank cells
"""
for index, cell in enumerate(nb.cells):
if cell.cell_type == 'code' and not cell.source:
nb.cells.pop(index)
else:
nb.cells[index], resources = self.preprocess_cell(cell, resources, index)
return nb, resources
def preprocess_cell(self, cell, resources, cell_index):
"""
Remove extraneous whitespace from code cells' source code
"""
if cell.cell_type == 'code':
cell.source = cell.source.strip()
return cell, resources
def doctor(string: str) -> str:
"""Get rid of all the wacky newlines nbconvert adds to markdown output and return result."""
post_code_newlines_patt = re.compile(r'(```)(\n+)')
inter_output_newlines_patt = re.compile(r'(\s{4}\S+)(\n+)(\s{4})')
post_code_filtered = re.sub(post_code_newlines_patt, r'\1\n\n', string)
inter_output_filtered = re.sub(inter_output_newlines_patt, r'\1\n\3', post_code_filtered)
return inter_output_filtered
def make_yaml_header(**kwargs):
header = '---\n'
for key, value in kwargs.items():
if type(value) in [str, int , float ] :
header += '{} : {}\n'.format(key, value)
else:
header += '{}: \n'.format(key)
for item in value:
header += ' - {}\n'.format(item)
header += '---\n'
return header
def notebook_to_markdown( path, date, slug, **kwargs ):
"""
Convert notebook to Markdown format
Args:
path: str, path to notebook
date: datestring in YYYY-MM-DD format
slug: str, front-matter parameter, used to compose adress of blogpost
kwargs: str, float, int, list, tuple, other front-matter parameters recommended to pass title
"""
path_nb = Path(path)
path_out = path_nb.parents[1] / 'static'/ date.split('-')[0] / date.split('-')[1] / slug
path_post = path_nb.parents[1] / 'content/post/' / ( date + '-' + slug + '.md' )
assert path_nb.exists()
assert path_post.parent.exists()
assert bool( re.match('[0-9]{4}-[0-1][0-9]-[0-3][0-9]', date) ), 'Incorrect date format, need YYYY-MM-DD'
# convert notebook to .md----------------------------------------------------
with Path(path).open() as fp:
notebook = nbformat.read(fp, as_version=4)
c = Config()
c.MarkdownExporter.preprocessors = [CustomPreprocessor]
markdown_exporter = MarkdownExporter(config=c)
markdown, resources = markdown_exporter.from_notebook_node(notebook)
md = doctor(markdown)
yaml = make_yaml_header( date = date
, slug = slug
, **kwargs)
md = yaml + md
with path_post.open('w') as f:
f.write(md)
# write outputs as png --------------------------------------------------------
if 'outputs' in resources.keys():
if not path_out.exists():
path_out.mkdir(parents=True)
for key in resources['outputs'].keys():
with (path_out / key).open('wb') as f:
f.write( resources['outputs'][key] )
if __name__ == "__main__":
path = r'../notebooks/adding_posts_with_juypter.ipynb'
notebook_to_markdown( path = r'../notebooks/adding_posts_with_juypter.ipynb'
, date = '2018-08-11'
, slug = 'post_with_jupyter'
, title = 'Adding posts as jupyter notebooks'
, author = 'Bjoern Koneswarakantha'
, categories = ['R vs. python','blogging']
, tags = ['R vs. python','hugo', 'python', 'R', 'hugo', 'hugo_jupyer']
, summary = 'Blogging with jupyter notebooks, hugo_jupyter and some tweaking. Comparison to\
R and blogdown'
, thumbnailImagePosition = 'left'
, thumbnailImage = 'https://gitlab.eurecom.fr/zoe-apps/pytorch/avatar'
)
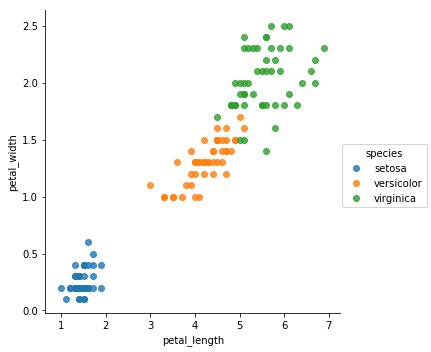
POC Graphical output
from matplotlib import pyplot as plt
%matplotlib inline
import seaborn as sns
df = sns.load_dataset('iris')
sns.lmplot(x = 'petal_length', y = 'petal_width', data = df
, hue = 'species'
, fit_reg = False)
<seaborn.axisgrid.FacetGrid at 0x251a9439518>
POC shortcode
Look we can use shortcode to wrap text in tags
